What’s New¶
v0.8.2 (18 August 2016)¶
This release includes a number of bug fixes and minor enhancements.
Breaking changes¶
broadcast()andconcat()now auto-align inputs, usingjoin=outer. Previously, these functions raisedValueErrorfor non-aligned inputs. By Guido Imperiale.
Enhancements¶
- New documentation on Transitioning from pandas.Panel to xarray. By Maximilian Roos.
- New
DatasetandDataArraymethodsto_dict()andfrom_dict()to allow easy conversion between dictionaries and xarray objects (GH432). See dictionary IO for more details. By Julia Signell. - Added
excludeandindexesoptional parameters toalign(), andexcludeoptional parameter tobroadcast(). By Guido Imperiale. - Better error message when assigning variables without dimensions (GH971). By Stephan Hoyer.
- Better error message when reindex/align fails due to duplicate index values (GH956). By Stephan Hoyer.
Bug fixes¶
- Ensure xarray works with h5netcdf v0.3.0 for arrays with
dtype=str(GH953). By Stephan Hoyer. Dataset.__dir__()(i.e. the method python calls to get autocomplete options) failed if one of the dataset’s keys was not a string (GH852). By Maximilian Roos.Datasetconstructor can now take arbitrary objects as values (GH647). By Maximilian Roos.- Clarified
copyargument forreindex()andalign(), which now consistently always return new xarray objects (GH927). - Fix
open_mfdatasetwithengine='pynio'(GH936). By Stephan Hoyer. groupby_binssorted bin labels as strings (GH952). By Stephan Hoyer.- Fix bug introduced by v0.8.0 that broke assignment to datasets when both the left and right side have the same non-unique index values (GH956).
v0.8.1 (5 August 2016)¶
Bug fixes¶
- Fix bug in v0.8.0 that broke assignment to Datasets with non-unique indexes (GH943). By Stephan Hoyer.
v0.8.0 (2 August 2016)¶
This release includes four months of new features and bug fixes, including several breaking changes.
Breaking changes¶
- Dropped support for Python 2.6 (GH855).
- Indexing on multi-index now drop levels, which is consistent with pandas. It also changes the name of the dimension / coordinate when the multi-index is reduced to a single index (GH802).
- Contour plots no longer add a colorbar per default (GH866). Filled contour plots are unchanged.
DataArray.valuesand.datanow always returns an NumPy array-like object, even for 0-dimensional arrays with object dtype (GH867). Previously,.valuesreturned native Python objects in such cases. To convert the values of scalar arrays to Python objects, use the.item()method.
Enhancements¶
- Groupby operations now support grouping over multidimensional variables. A new
method called
groupby_bins()has also been added to allow users to specify bins for grouping. The new features are described in Multidimensional Grouping and Working with Multidimensional Coordinates. By Ryan Abernathey. - DataArray and Dataset method
where()now supports adrop=Trueoption that clips coordinate elements that are fully masked. By Phillip J. Wolfram. - New top level
merge()function allows for combining variables from any number ofDatasetand/orDataArrayvariables. See Merge for more details. By Stephan Hoyer. - DataArray and Dataset method
resample()now supports thekeep_attrs=Falseoption that determines whether variable and dataset attributes are retained in the resampled object. By Jeremy McGibbon. - Better multi-index support in DataArray and Dataset
sel()andloc()methods, which now behave more closely to pandas and which also accept dictionaries for indexing based on given level names and labels (see Multi-level indexing). By Benoit Bovy. - New (experimental) decorators
register_dataset_accessor()andregister_dataarray_accessor()for registering custom xarray extensions without subclassing. They are described in the new documentation page on xarray Internals. By Stephan Hoyer. - Round trip boolean datatypes. Previously, writing boolean datatypes to netCDF formats would raise an error since netCDF does not have a bool datatype. This feature reads/writes a dtype attribute to boolean variables in netCDF files. By Joe Hamman.
- 2D plotting methods now have two new keywords (cbar_ax and cbar_kwargs), allowing more control on the colorbar (GH872). By Fabien Maussion.
- New Dataset method
filter_by_attrs(), akin tonetCDF4.Dataset.get_variables_by_attributes, to easily filter data variables using its attributes. Filipe Fernandes.
Bug fixes¶
Attributes were being retained by default for some resampling operations when they should not. With the
keep_attrs=Falseoption, they will no longer be retained by default. This may be backwards-incompatible with some scripts, but the attributes may be kept by adding thekeep_attrs=Trueoption. By Jeremy McGibbon.Concatenating xarray objects along an axis with a MultiIndex or PeriodIndex preserves the nature of the index (GH875). By Stephan Hoyer.
Fixed bug in arithmetic operations on DataArray objects whose dimensions are numpy structured arrays or recarrays GH861, GH837. By Maciek Swat.
decode_cf_timedeltanow accepts arrays withndim>1 (GH842).This fixes issue GH665. Filipe Fernandes.
Fix a bug where xarray.ufuncs that take two arguments would incorrectly use to numpy functions instead of dask.array functions (GH876). By Stephan Hoyer.
Support for pickling functions from
xarray.ufuncs(GH901). By Stephan Hoyer.Variable.copy(deep=True)no longer converts MultiIndex into a base Index (GH769). By Benoit Bovy.Fixes for groupby on dimensions with a multi-index (GH867). By Stephan Hoyer.
Fix printing datasets with unicode attributes on Python 2 (GH892). By Stephan Hoyer.
Fixed incorrect test for dask version (GH891). By Stephan Hoyer.
Fixed dim argument for isel_points/sel_points when a pandas.Index is passed. By Stephan Hoyer.
contour()now plots the correct number of contours (GH866). By Fabien Maussion.
v0.7.2 (13 March 2016)¶
This release includes two new, entirely backwards compatible features and several bug fixes.
Enhancements¶
New DataArray method
DataArray.dot()for calculating the dot product of two DataArrays along shared dimensions. By Dean Pospisil.Rolling window operations on DataArray objects are now supported via a new
DataArray.rolling()method. For example:In [1]: import xarray as xr; import numpy as np In [2]: arr = xr.DataArray(np.arange(0, 7.5, 0.5).reshape(3, 5), dims=('x', 'y')) In [3]: arr Out[3]: <xarray.DataArray (x: 3, y: 5)> array([[ 0. , 0.5, 1. , 1.5, 2. ], [ 2.5, 3. , 3.5, 4. , 4.5], [ 5. , 5.5, 6. , 6.5, 7. ]]) Coordinates: * x (x) int64 0 1 2 * y (y) int64 0 1 2 3 4 In [4]: arr.rolling(y=3, min_periods=2).mean() Out[4]: <xarray.DataArray (x: 3, y: 5)> array([[ nan, 0.25, 0.5 , 1. , 1.5 ], [ nan, 2.75, 3. , 3.5 , 4. ], [ nan, 5.25, 5.5 , 6. , 6.5 ]]) Coordinates: * x (x) int64 0 1 2 * y (y) int64 0 1 2 3 4
See Rolling window operations for more details. By Joe Hamman.
Bug fixes¶
- Fixed an issue where plots using pcolormesh and Cartopy axes were being distorted
by the inference of the axis interval breaks. This change chooses not to modify
the coordinate variables when the axes have the attribute
projection, allowing Cartopy to handle the extent of pcolormesh plots (GH781). By Joe Hamman. - 2D plots now better handle additional coordinates which are not
DataArraydimensions (GH788). By Fabien Maussion.
v0.7.1 (16 February 2016)¶
This is a bug fix release that includes two small, backwards compatible enhancements. We recommend that all users upgrade.
Enhancements¶
Bug fixes¶
- Restore checks for shape consistency between data and coordinates in the DataArray constructor (GH758).
- Single dimension variables no longer transpose as part of a broader
.transpose. This behavior was causingpandas.PeriodIndexdimensions to lose their type (GH749) Datasetlabels remain as their native type on.to_dataset. Previously they were coerced to strings (GH745)- Fixed a bug where replacing a
DataArrayindex coordinate would improperly align the coordinate (GH725). DataArray.reindex_likenow maintains the dtype of complex numbers when reindexing leads to NaN values (GH738).Dataset.renameandDataArray.renamesupport the old and new names being the same (GH724).- Fix
from_dataset()for DataFrames with Categorical column and a MultiIndex index (GH737). - Fixes to ensure xarray works properly after the upcoming pandas v0.18 and NumPy v1.11 releases.
Acknowledgments¶
The following individuals contributed to this release:
- Edward Richards
- Maximilian Roos
- Rafael Guedes
- Spencer Hill
- Stephan Hoyer
v0.7.0 (21 January 2016)¶
This major release includes redesign of DataArray
internals, as well as new methods for reshaping, rolling and shifting
data. It includes preliminary support for pandas.MultiIndex,
as well as a number of other features and bug fixes, several of which
offer improved compatibility with pandas.
New name¶
The project formerly known as “xray” is now “xarray”, pronounced “x-array”! This avoids a namespace conflict with the entire field of x-ray science. Renaming our project seemed like the right thing to do, especially because some scientists who work with actual x-rays are interested in using this project in their work. Thanks for your understanding and patience in this transition. You can now find our documentation and code repository at new URLs:
To ease the transition, we have simultaneously released v0.7.0 of both
xray and xarray on the Python Package Index. These packages are
identical. For now, import xray still works, except it issues a
deprecation warning. This will be the last xray release. Going forward, we
recommend switching your import statements to import xarray as xr.
Breaking changes¶
The internal data model used by
DataArrayhas been rewritten to fix several outstanding issues (GH367, GH634, this stackoverflow report). Internally,DataArrayis now implemented in terms of._variableand._coordsattributes instead of holding variables in aDatasetobject.This refactor ensures that if a DataArray has the same name as one of its coordinates, the array and the coordinate no longer share the same data.
In practice, this means that creating a DataArray with the same
nameas one of its dimensions no longer automatically uses that array to label the corresponding coordinate. You will now need to provide coordinate labels explicitly. Here’s the old behavior:In [5]: xray.DataArray([4, 5, 6], dims='x', name='x') Out[5]: <xray.DataArray 'x' (x: 3)> array([4, 5, 6]) Coordinates: * x (x) int64 4 5 6
and the new behavior (compare the values of the
xcoordinate):In [6]: xray.DataArray([4, 5, 6], dims='x', name='x') Out[6]: <xray.DataArray 'x' (x: 3)> array([4, 5, 6]) Coordinates: * x (x) int64 0 1 2
It is no longer possible to convert a DataArray to a Dataset with
xray.DataArray.to_dataset()if it is unnamed. This will now raiseValueError. If the array is unnamed, you need to supply thenameargument.
Enhancements¶
Basic support for
MultiIndexcoordinates on xray objects, including indexing,stack()andunstack():In [7]: df = pd.DataFrame({'foo': range(3), ...: 'x': ['a', 'b', 'b'], ...: 'y': [0, 0, 1]}) ...: In [8]: s = df.set_index(['x', 'y'])['foo'] In [9]: arr = xray.DataArray(s, dims='z') In [10]: arr Out[10]: <xray.DataArray 'foo' (z: 3)> array([0, 1, 2]) Coordinates: * z (z) object ('a', 0) ('b', 0) ('b', 1) In [11]: arr.indexes['z'] Out[11]: MultiIndex(levels=[[u'a', u'b'], [0, 1]], labels=[[0, 1, 1], [0, 0, 1]], names=[u'x', u'y']) In [12]: arr.unstack('z') Out[12]: <xray.DataArray 'foo' (x: 2, y: 2)> array([[ 0., nan], [ 1., 2.]]) Coordinates: * x (x) object 'a' 'b' * y (y) int64 0 1 In [13]: arr.unstack('z').stack(z=('x', 'y')) Out[13]: <xray.DataArray 'foo' (z: 4)> array([ 0., nan, 1., 2.]) Coordinates: * z (z) object ('a', 0) ('a', 1) ('b', 0) ('b', 1)
See Stack and unstack for more details.
Warning
xray’s MultiIndex support is still experimental, and we have a long to- do list of desired additions (GH719), including better display of multi-index levels when printing a
Dataset, and support for saving datasets with a MultiIndex to a netCDF file. User contributions in this area would be greatly appreciated.Support for reading GRIB, HDF4 and other file formats via PyNIO. See Formats supported by PyNIO for more details.
Better error message when a variable is supplied with the same name as one of its dimensions.
Plotting: more control on colormap parameters (GH642).
vminandvmaxwill not be silently ignored anymore. Settingcenter=Falseprevents automatic selection of a divergent colormap.New
shift()androll()methods for shifting/rotating datasets or arrays along a dimension:In [14]: array = xray.DataArray([5, 6, 7, 8], dims='x') In [15]: array.shift(x=2) Out[15]: <xarray.DataArray (x: 4)> array([ nan, nan, 5., 6.]) Coordinates: * x (x) int64 0 1 2 3 In [16]: array.roll(x=2) Out[16]: <xarray.DataArray (x: 4)> array([7, 8, 5, 6]) Coordinates: * x (x) int64 2 3 0 1
Notice that
shiftmoves data independently of coordinates, butrollmoves both data and coordinates.Assigning a
pandasobject directly as aDatasetvariable is now permitted. Its index names correspond to thedimsof theDataset, and its data is aligned.Passing a
pandas.DataFrameorpandas.Panelto a Dataset constructor is now permitted.New function
broadcast()for explicitly broadcastingDataArrayandDatasetobjects against each other. For example:In [17]: a = xray.DataArray([1, 2, 3], dims='x') In [18]: b = xray.DataArray([5, 6], dims='y') In [19]: a Out[19]: <xarray.DataArray (x: 3)> array([1, 2, 3]) Coordinates: * x (x) int64 0 1 2 In [20]: b Out[20]: <xarray.DataArray (y: 2)> array([5, 6]) Coordinates: * y (y) int64 0 1 In [21]: a2, b2 = xray.broadcast(a, b) In [22]: a2 Out[22]: <xarray.DataArray (x: 3, y: 2)> array([[1, 1], [2, 2], [3, 3]]) Coordinates: * x (x) int64 0 1 2 * y (y) int64 0 1 In [23]: b2 Out[23]: <xarray.DataArray (x: 3, y: 2)> array([[5, 6], [5, 6], [5, 6]]) Coordinates: * y (y) int64 0 1 * x (x) int64 0 1 2
Bug fixes¶
- Fixes for several issues found on
DataArrayobjects with the same name as one of their coordinates (see Breaking changes for more details). DataArray.to_masked_arrayalways returns masked array with mask being an array (not a scalar value) (GH684)- Allows for (imperfect) repr of Coords when underlying index is PeriodIndex (GH645).
- Fixes for several issues found on
DataArrayobjects with the same name as one of their coordinates (see Breaking changes for more details). - Attempting to assign a
DatasetorDataArrayvariable/attribute using attribute-style syntax (e.g.,ds.foo = 42) now raises an error rather than silently failing (GH656, GH714). - You can now pass pandas objects with non-numpy dtypes (e.g.,
categoricalordatetime64with a timezone) into xray without an error (GH716).
Acknowledgments¶
The following individuals contributed to this release:
- Antony Lee
- Fabien Maussion
- Joe Hamman
- Maximilian Roos
- Stephan Hoyer
- Takeshi Kanmae
- femtotrader
v0.6.1 (21 October 2015)¶
This release contains a number of bug and compatibility fixes, as well as enhancements to plotting, indexing and writing files to disk.
Note that the minimum required version of dask for use with xray is now version 0.6.
API Changes¶
- The handling of colormaps and discrete color lists for 2D plots in
plot()was changed to provide more compatibility with matplotlib’scontourandcontourffunctions (GH538). Now discrete lists of colors should be specified usingcolorskeyword, rather thancmap.
Enhancements¶
Faceted plotting through
FacetGridand theplot()method. See Faceting for more details and examples.sel()andreindex()now support thetoleranceargument for controlling nearest-neighbor selection (GH629):In [24]: array = xray.DataArray([1, 2, 3], dims='x') In [25]: array.reindex(x=[0.9, 1.5], method='nearest', tolerance=0.2) Out[25]: <xray.DataArray (x: 2)> array([ 2., nan]) Coordinates: * x (x) float64 0.9 1.5
This feature requires pandas v0.17 or newer.
New
encodingargument into_netcdf()for writing netCDF files with compression, as described in the new documentation section on Writing encoded data.Add
realandimagattributes to Dataset and DataArray (GH553).More informative error message with
from_dataframe()if the frame has duplicate columns.xray now uses deterministic names for dask arrays it creates or opens from disk. This allows xray users to take advantage of dask’s nascent support for caching intermediate computation results. See GH555 for an example.
Bug fixes¶
- Forwards compatibility with the latest pandas release (v0.17.0). We were using some internal pandas routines for datetime conversion, which unfortunately have now changed upstream (GH569).
- Aggregation functions now correctly skip
NaNfor data forcomplex128dtype (GH554). - Fixed indexing 0d arrays with unicode dtype (GH568).
name()and Dataset keys must be a string or None to be written to netCDF (GH533).where()now uses dask instead of numpy if either the array orotheris a dask array. Previously, ifotherwas a numpy array the method was evaluated eagerly.- Global attributes are now handled more consistently when loading remote
datasets using
engine='pydap'(GH574). - It is now possible to assign to the
.dataattribute of DataArray objects. coordinatesattribute is now kept in the encoding dictionary after decoding (GH610).- Compatibility with numpy 1.10 (GH617).
Acknowledgments¶
The following individuals contributed to this release:
- Ryan Abernathey
- Pete Cable
- Clark Fitzgerald
- Joe Hamman
- Stephan Hoyer
- Scott Sinclair
v0.6.0 (21 August 2015)¶
This release includes numerous bug fixes and enhancements. Highlights
include the introduction of a plotting module and the new Dataset and DataArray
methods isel_points(), sel_points(),
where() and diff(). There are no
breaking changes from v0.5.2.
Enhancements¶
Plotting methods have been implemented on DataArray objects
plot()through integration with matplotlib (GH185). For an introduction, see Plotting.Variables in netCDF files with multiple missing values are now decoded as NaN after issuing a warning if open_dataset is called with mask_and_scale=True.
We clarified our rules for when the result from an xray operation is a copy vs. a view (see Copies vs. views for more details).
Dataset variables are now written to netCDF files in order of appearance when using the netcdf4 backend (GH479).
Added
isel_points()andsel_points()to support pointwise indexing of Datasets and DataArrays (GH475).In [26]: da = xray.DataArray(np.arange(56).reshape((7, 8)), ....: coords={'x': list('abcdefg'), ....: 'y': 10 * np.arange(8)}, ....: dims=['x', 'y']) ....: In [27]: da Out[27]: <xray.DataArray (x: 7, y: 8)> array([[ 0, 1, 2, 3, 4, 5, 6, 7], [ 8, 9, 10, 11, 12, 13, 14, 15], [16, 17, 18, 19, 20, 21, 22, 23], [24, 25, 26, 27, 28, 29, 30, 31], [32, 33, 34, 35, 36, 37, 38, 39], [40, 41, 42, 43, 44, 45, 46, 47], [48, 49, 50, 51, 52, 53, 54, 55]]) Coordinates: * y (y) int64 0 10 20 30 40 50 60 70 * x (x) |S1 'a' 'b' 'c' 'd' 'e' 'f' 'g' # we can index by position along each dimension In [28]: da.isel_points(x=[0, 1, 6], y=[0, 1, 0], dim='points') Out[28]: <xray.DataArray (points: 3)> array([ 0, 9, 48]) Coordinates: y (points) int64 0 10 0 x (points) |S1 'a' 'b' 'g' * points (points) int64 0 1 2 # or equivalently by label In [29]: da.sel_points(x=['a', 'b', 'g'], y=[0, 10, 0], dim='points') Out[29]: <xray.DataArray (points: 3)> array([ 0, 9, 48]) Coordinates: y (points) int64 0 10 0 x (points) |S1 'a' 'b' 'g' * points (points) int64 0 1 2
New
where()method for masking xray objects according to some criteria. This works particularly well with multi-dimensional data:In [30]: ds = xray.Dataset(coords={'x': range(100), 'y': range(100)}) In [31]: ds['distance'] = np.sqrt(ds.x ** 2 + ds.y ** 2) In [32]: ds.distance.where(ds.distance < 100).plot() Out[32]: <matplotlib.collections.QuadMesh at 0x7f07d33f8790>
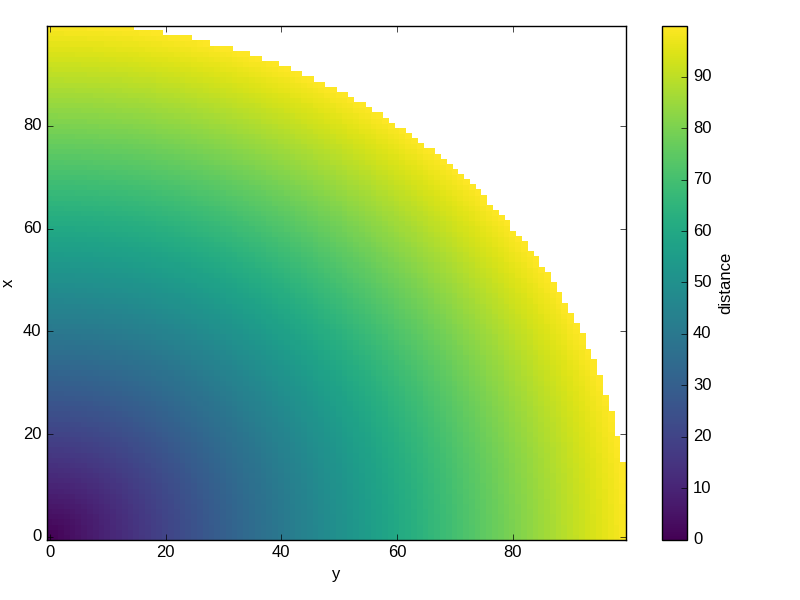
Added new methods
DataArray.diffandDataset.difffor finite difference calculations along a given axis.New
to_masked_array()convenience method for returning a numpy.ma.MaskedArray.In [33]: da = xray.DataArray(np.random.random_sample(size=(5, 4))) In [34]: da.where(da < 0.5) Out[34]: <xarray.DataArray (dim_0: 5, dim_1: 4)> array([[ 0.127, nan, 0.26 , nan], [ 0.377, 0.336, 0.451, nan], [ 0.123, nan, 0.373, 0.448], [ 0.129, nan, nan, 0.352], [ 0.229, nan, nan, 0.138]]) Coordinates: * dim_0 (dim_0) int64 0 1 2 3 4 * dim_1 (dim_1) int64 0 1 2 3 In [35]: da.where(da < 0.5).to_masked_array(copy=True) Out[35]: masked_array(data = [[0.12696983303810094 -- 0.26047600586578334 --] [0.37674971618967135 0.33622174433445307 0.45137647047539964 --] [0.12310214428849964 -- 0.37301222522143085 0.4479968246859435] [0.12944067971751294 -- -- 0.35205353914802473] [0.2288873043216132 -- -- 0.1375535565632705]], mask = [[False True False True] [False False False True] [False True False False] [False True True False] [False True True False]], fill_value = 1e+20)
Added new flag “drop_variables” to
open_dataset()for excluding variables from being parsed. This may be useful to drop variables with problems or inconsistent values.
Bug fixes¶
- Fixed aggregation functions (e.g., sum and mean) on big-endian arrays when bottleneck is installed (GH489).
- Dataset aggregation functions dropped variables with unsigned integer dtype (GH505).
.any()and.all()were not lazy when used on xray objects containing dask arrays.- Fixed an error when attempting to saving datetime64 variables to netCDF
files when the first element is
NaT(GH528). - Fix pickle on DataArray objects (GH515).
- Fixed unnecessary coercion of float64 to float32 when using netcdf3 and netcdf4_classic formats (GH526).
v0.5.2 (16 July 2015)¶
This release contains bug fixes, several additional options for opening and
saving netCDF files, and a backwards incompatible rewrite of the advanced
options for xray.concat.
Backwards incompatible changes¶
- The optional arguments
concat_overandmodeinconcat()have been removed and replaced bydata_varsandcoords. The new arguments are both more easily understood and more robustly implemented, and allowed us to fix a bug whereconcataccidentally loaded data into memory. If you set values for these optional arguments manually, you will need to update your code. The default behavior should be unchanged.
Enhancements¶
open_mfdataset()now supports apreprocessargument for preprocessing datasets prior to concatenaton. This is useful if datasets cannot be otherwise merged automatically, e.g., if the original datasets have conflicting index coordinates (GH443).open_dataset()andopen_mfdataset()now use a global thread lock by default for reading from netCDF files with dask. This avoids possible segmentation faults for reading from netCDF4 files when HDF5 is not configured properly for concurrent access (GH444).Added support for serializing arrays of complex numbers with engine=’h5netcdf’.
The new
save_mfdataset()function allows for saving multiple datasets to disk simultaneously. This is useful when processing large datasets with dask.array. For example, to save a dataset too big to fit into memory to one file per year, we could write:In [36]: years, datasets = zip(*ds.groupby('time.year')) In [37]: paths = ['%s.nc' % y for y in years] In [38]: xray.save_mfdataset(datasets, paths)
Bug fixes¶
- Fixed
min,max,argminandargmaxfor arrays with string or unicode types (GH453). open_dataset()andopen_mfdataset()support supplying chunks as a single integer.- Fixed a bug in serializing scalar datetime variable to netCDF.
- Fixed a bug that could occur in serialization of 0-dimensional integer arrays.
- Fixed a bug where concatenating DataArrays was not always lazy (GH464).
- When reading datasets with h5netcdf, bytes attributes are decoded to strings. This allows conventions decoding to work properly on Python 3 (GH451).
v0.5.1 (15 June 2015)¶
This minor release fixes a few bugs and an inconsistency with pandas. It also
adds the pipe method, copied from pandas.
Enhancements¶
- Added
pipe(), replicating the new pandas method in version 0.16.2. See Transforming datasets for more details. assign()andassign_coords()now assign new variables in sorted (alphabetical) order, mirroring the behavior in pandas. Previously, the order was arbitrary.
v0.5 (1 June 2015)¶
Highlights¶
The headline feature in this release is experimental support for out-of-core
computing (data that doesn’t fit into memory) with dask. This includes a new
top-level function open_mfdataset() that makes it easy to open
a collection of netCDF (using dask) as a single xray.Dataset object. For
more on dask, read the blog post introducing xray + dask and the new
documentation section Out of core computation with dask.
Dask makes it possible to harness parallelism and manipulate gigantic datasets with xray. It is currently an optional dependency, but it may become required in the future.
Backwards incompatible changes¶
The logic used for choosing which variables are concatenated with
concat()has changed. Previously, by default any variables which were equal across a dimension were not concatenated. This lead to some surprising behavior, where the behavior of groupby and concat operations could depend on runtime values (GH268). For example:In [39]: ds = xray.Dataset({'x': 0}) In [40]: xray.concat([ds, ds], dim='y') Out[40]: <xray.Dataset> Dimensions: () Coordinates: *empty* Data variables: x int64 0
Now, the default always concatenates data variables:
In [41]: xray.concat([ds, ds], dim='y') Out[41]: <xarray.Dataset> Dimensions: (y: 2) Coordinates: * y (y) int64 0 1 Data variables: x (y) int64 0 0
To obtain the old behavior, supply the argument
concat_over=[].
Enhancements¶
New
to_array()and enhancedto_dataset()methods make it easy to switch back and forth between arrays and datasets:In [42]: ds = xray.Dataset({'a': 1, 'b': ('x', [1, 2, 3])}, ....: coords={'c': 42}, attrs={'Conventions': 'None'}) ....: In [43]: ds.to_array() Out[43]: <xarray.DataArray (variable: 2, x: 3)> array([[1, 1, 1], [1, 2, 3]]) Coordinates: * variable (variable) |S1 'a' 'b' * x (x) int64 0 1 2 c int64 42 Attributes: Conventions: None In [44]: ds.to_array().to_dataset(dim='variable') Out[44]: <xarray.Dataset> Dimensions: (x: 3) Coordinates: * x (x) int64 0 1 2 c int64 42 Data variables: a (x) int64 1 1 1 b (x) int64 1 2 3 Attributes: Conventions: None
New
fillna()method to fill missing values, modeled off the pandas method of the same name:In [45]: array = xray.DataArray([np.nan, 1, np.nan, 3], dims='x') In [46]: array.fillna(0) Out[46]: <xarray.DataArray (x: 4)> array([ 0., 1., 0., 3.]) Coordinates: * x (x) int64 0 1 2 3
fillnaworks on bothDatasetandDataArrayobjects, and uses index based alignment and broadcasting like standard binary operations. It also can be applied by group, as illustrated in Fill missing values with climatology.New
assign()andassign_coords()methods patterned off the newDataFrame.assignmethod in pandas:In [47]: ds = xray.Dataset({'y': ('x', [1, 2, 3])}) In [48]: ds.assign(z = lambda ds: ds.y ** 2) Out[48]: <xarray.Dataset> Dimensions: (x: 3) Coordinates: * x (x) int64 0 1 2 Data variables: y (x) int64 1 2 3 z (x) int64 1 4 9 In [49]: ds.assign_coords(z = ('x', ['a', 'b', 'c'])) Out[49]: <xarray.Dataset> Dimensions: (x: 3) Coordinates: * x (x) int64 0 1 2 z (x) |S1 'a' 'b' 'c' Data variables: y (x) int64 1 2 3
These methods return a new Dataset (or DataArray) with updated data or coordinate variables.
sel()now supports themethodparameter, which works like the paramter of the same name onreindex(). It provides a simple interface for doing nearest-neighbor interpolation:In [50]: ds.sel(x=1.1, method='nearest') Out[50]: <xray.Dataset> Dimensions: () Coordinates: x int64 1 Data variables: y int64 2 In [51]: ds.sel(x=[1.1, 2.1], method='pad') Out[51]: <xray.Dataset> Dimensions: (x: 2) Coordinates: * x (x) int64 1 2 Data variables: y (x) int64 2 3
See Nearest neighbor lookups for more details.
You can now control the underlying backend used for accessing remote datasets (via OPeNDAP) by specifying
engine='netcdf4'orengine='pydap'.xray now provides experimental support for reading and writing netCDF4 files directly via h5py with the h5netcdf package, avoiding the netCDF4-Python package. You will need to install h5netcdf and specify
engine='h5netcdf'to try this feature.Accessing data from remote datasets now has retrying logic (with exponential backoff) that should make it robust to occasional bad responses from DAP servers.
You can control the width of the Dataset repr with
xray.set_options. It can be used either as a context manager, in which case the default is restored outside the context:In [52]: ds = xray.Dataset({'x': np.arange(1000)}) In [53]: with xray.set_options(display_width=40): ....: print(ds) ....: <xarray.Dataset> Dimensions: (x: 1000) Coordinates: * x (x) int64 0 1 2 3 4 5 6 ... Data variables: *empty*
Or to set a global option:
In [54]: xray.set_options(display_width=80)
The default value for the
display_widthoption is 80.
Deprecations¶
- The method
load_data()has been renamed to the more succinctload().
v0.4.1 (18 March 2015)¶
The release contains bug fixes and several new features. All changes should be fully backwards compatible.
Enhancements¶
New documentation sections on Time series data and Combining multiple files.
resample()lets you resample a dataset or data array to a new temporal resolution. The syntax is the same as pandas, except you need to supply the time dimension explicitly:In [55]: time = pd.date_range('2000-01-01', freq='6H', periods=10) In [56]: array = xray.DataArray(np.arange(10), [('time', time)]) In [57]: array.resample('1D', dim='time') Out[57]: <xarray.DataArray (time: 3)> array([ 1.5, 5.5, 8.5]) Coordinates: * time (time) datetime64[ns] 2000-01-01 2000-01-02 2000-01-03
You can specify how to do the resampling with the
howargument and other options such asclosedandlabellet you control labeling:In [58]: array.resample('1D', dim='time', how='sum', label='right') Out[58]: <xarray.DataArray (time: 3)> array([ 6, 22, 17]) Coordinates: * time (time) datetime64[ns] 2000-01-02 2000-01-03 2000-01-04
If the desired temporal resolution is higher than the original data (upsampling), xray will insert missing values:
In [59]: array.resample('3H', 'time') Out[59]: <xarray.DataArray (time: 19)> array([ 0., nan, 1., ..., 8., nan, 9.]) Coordinates: * time (time) datetime64[ns] 2000-01-01 2000-01-01T03:00:00 ...
firstandlastmethods on groupby objects let you take the first or last examples from each group along the grouped axis:In [60]: array.groupby('time.day').first() Out[60]: <xarray.DataArray (day: 3)> array([0, 4, 8]) Coordinates: * day (day) int64 1 2 3
These methods combine well with
resample:In [61]: array.resample('1D', dim='time', how='first') Out[61]: <xarray.DataArray (time: 3)> array([0, 4, 8]) Coordinates: * time (time) datetime64[ns] 2000-01-01 2000-01-02 2000-01-03
swap_dims()allows for easily swapping one dimension out for another:In [62]: ds = xray.Dataset({'x': range(3), 'y': ('x', list('abc'))}) In [63]: ds Out[63]: <xarray.Dataset> Dimensions: (x: 3) Coordinates: * x (x) int64 0 1 2 Data variables: y (x) |S1 'a' 'b' 'c' In [64]: ds.swap_dims({'x': 'y'}) Out[64]: <xarray.Dataset> Dimensions: (y: 3) Coordinates: * y (y) |S1 'a' 'b' 'c' x (y) int64 0 1 2 Data variables: *empty*
This was possible in earlier versions of xray, but required some contortions.
open_dataset()andto_netcdf()now accept anengineargument to explicitly select which underlying library (netcdf4 or scipy) is used for reading/writing a netCDF file.
Bug fixes¶
- Fixed a bug where data netCDF variables read from disk with
engine='scipy'could still be associated with the file on disk, even after closing the file (GH341). This manifested itself in warnings about mmapped arrays and segmentation faults (if the data was accessed). - Silenced spurious warnings about all-NaN slices when using nan-aware aggregation methods (GH344).
- Dataset aggregations with
keep_attrs=Truenow preserve attributes on data variables, not just the dataset itself. - Tests for xray now pass when run on Windows (GH360).
- Fixed a regression in v0.4 where saving to netCDF could fail with the error
ValueError: could not automatically determine time units.
v0.4 (2 March, 2015)¶
This is one of the biggest releases yet for xray: it includes some major changes that may break existing code, along with the usual collection of minor enhancements and bug fixes. On the plus side, this release includes all hitherto planned breaking changes, so the upgrade path for xray should be smoother going forward.
Breaking changes¶
We now automatically align index labels in arithmetic, dataset construction, merging and updating. This means the need for manually invoking methods like
align()andreindex_like()should be vastly reduced.For arithmetic, we align based on the intersection of labels:
In [65]: lhs = xray.DataArray([1, 2, 3], [('x', [0, 1, 2])]) In [66]: rhs = xray.DataArray([2, 3, 4], [('x', [1, 2, 3])]) In [67]: lhs + rhs Out[67]: <xarray.DataArray (x: 2)> array([4, 6]) Coordinates: * x (x) int64 1 2
For dataset construction and merging, we align based on the union of labels:
In [68]: xray.Dataset({'foo': lhs, 'bar': rhs}) Out[68]: <xarray.Dataset> Dimensions: (x: 4) Coordinates: * x (x) int64 0 1 2 3 Data variables: foo (x) float64 1.0 2.0 3.0 nan bar (x) float64 nan 2.0 3.0 4.0
For update and __setitem__, we align based on the original object:
In [69]: lhs.coords['rhs'] = rhs In [70]: lhs Out[70]: <xarray.DataArray (x: 3)> array([1, 2, 3]) Coordinates: * x (x) int64 0 1 2 rhs (x) float64 nan 2.0 3.0
Aggregations like
meanormediannow skip missing values by default:In [71]: xray.DataArray([1, 2, np.nan, 3]).mean() Out[71]: <xarray.DataArray ()> array(2.0)
You can turn this behavior off by supplying the keyword arugment
skipna=False.These operations are lightning fast thanks to integration with bottleneck, which is a new optional dependency for xray (numpy is used if bottleneck is not installed).
Scalar coordinates no longer conflict with constant arrays with the same value (e.g., in arithmetic, merging datasets and concat), even if they have different shape (GH243). For example, the coordinate
chere persists through arithmetic, even though it has different shapes on each DataArray:In [72]: a = xray.DataArray([1, 2], coords={'c': 0}, dims='x') In [73]: b = xray.DataArray([1, 2], coords={'c': ('x', [0, 0])}, dims='x') In [74]: (a + b).coords Out[74]: Coordinates: c (x) int64 0 0 * x (x) int64 0 1
This functionality can be controlled through the
compatoption, which has also been added to theDatasetconstructor.Datetime shortcuts such as
'time.month'now return aDataArraywith the name'month', not'time.month'(GH345). This makes it easier to index the resulting arrays when they are used withgroupby:In [75]: time = xray.DataArray(pd.date_range('2000-01-01', periods=365), ....: dims='time', name='time') ....: In [76]: counts = time.groupby('time.month').count() In [77]: counts.sel(month=2) Out[77]: <xarray.DataArray 'time' ()> array(29) Coordinates: month int64 2
Previously, you would need to use something like
counts.sel(**{'time.month': 2}}), which is much more awkward.The
seasondatetime shortcut now returns an array of string labels such ‘DJF’:In [78]: ds = xray.Dataset({'t': pd.date_range('2000-01-01', periods=12, freq='M')}) In [79]: ds['t.season'] Out[79]: <xarray.DataArray 'season' (t: 12)> array(['DJF', 'DJF', 'MAM', ..., 'SON', 'SON', 'DJF'], dtype='|S3') Coordinates: * t (t) datetime64[ns] 2000-01-31 2000-02-29 2000-03-31 2000-04-30 ...
Previously, it returned numbered seasons 1 through 4.
We have updated our use of the terms of “coordinates” and “variables”. What were known in previous versions of xray as “coordinates” and “variables” are now referred to throughout the documentation as “coordinate variables” and “data variables”. This brings xray in closer alignment to CF Conventions. The only visible change besides the documentation is that
Dataset.varshas been renamedDataset.data_vars.You will need to update your code if you have been ignoring deprecation warnings: methods and attributes that were deprecated in xray v0.3 or earlier (e.g.,
dimensions,attributes`) have gone away.
Enhancements¶
Support for
reindex()with a fill method. This provides a useful shortcut for upsampling:In [80]: data = xray.DataArray([1, 2, 3], dims='x') In [81]: data.reindex(x=[0.5, 1, 1.5, 2, 2.5], method='pad') Out[81]: <xarray.DataArray (x: 5)> array([1, 2, 2, 3, 3]) Coordinates: * x (x) float64 0.5 1.0 1.5 2.0 2.5
This will be especially useful once pandas 0.16 is released, at which point xray will immediately support reindexing with method=’nearest’.
Use functions that return generic ndarrays with DataArray.groupby.apply and Dataset.apply (GH327 and GH329). Thanks Jeff Gerard!
Consolidated the functionality of
dumps(writing a dataset to a netCDF3 bytestring) intoto_netcdf()(GH333).to_netcdf()now supports writing to groups in netCDF4 files (GH333). It also finally has a full docstring – you should read it!open_dataset()andto_netcdf()now work on netCDF3 files when netcdf4-python is not installed as long as scipy is available (GH333).The new
Dataset.dropandDataArray.dropmethods makes it easy to drop explicitly listed variables or index labels:# drop variables In [82]: ds = xray.Dataset({'x': 0, 'y': 1}) In [83]: ds.drop('x') Out[83]: <xarray.Dataset> Dimensions: () Coordinates: *empty* Data variables: y int64 1 # drop index labels In [84]: arr = xray.DataArray([1, 2, 3], coords=[('x', list('abc'))]) In [85]: arr.drop(['a', 'c'], dim='x') Out[85]: <xarray.DataArray (x: 1)> array([2]) Coordinates: * x (x) |S1 'b'
broadcast_equals()has been added to correspond to the newcompatoption.Long attributes are now truncated at 500 characters when printing a dataset (GH338). This should make things more convenient for working with datasets interactively.
Added a new documentation example, Calculating Seasonal Averages from Timeseries of Monthly Means. Thanks Joe Hamman!
Bug fixes¶
- Several bug fixes related to decoding time units from netCDF files (GH316, GH330). Thanks Stefan Pfenninger!
- xray no longer requires
decode_coords=Falsewhen reading datasets with unparseable coordinate attributes (GH308). - Fixed
DataArray.locindexing with...(GH318). - Fixed an edge case that resulting in an error when reindexing multi-dimensional variables (GH315).
- Slicing with negative step sizes (GH312).
- Invalid conversion of string arrays to numeric dtype (GH305).
- Fixed``repr()`` on dataset objects with non-standard dates (GH347).
Deprecations¶
dumpanddumpshave been deprecated in favor ofto_netcdf().drop_varshas been deprecated in favor ofdrop().
Future plans¶
The biggest feature I’m excited about working toward in the immediate future is supporting out-of-core operations in xray using Dask, a part of the Blaze project. For a preview of using Dask with weather data, read this blog post by Matthew Rocklin. See GH328 for more details.
v0.3.2 (23 December, 2014)¶
This release focused on bug-fixes, speedups and resolving some niggling inconsistencies.
There are a few cases where the behavior of xray differs from the previous version. However, I expect that in almost all cases your code will continue to run unmodified.
Warning
xray now requires pandas v0.15.0 or later. This was necessary for supporting TimedeltaIndex without too many painful hacks.
Backwards incompatible changes¶
Arrays of
datetime.datetimeobjects are now automatically cast todatetime64[ns]arrays when stored in an xray object, using machinery borrowed from pandas:In [86]: from datetime import datetime In [87]: xray.Dataset({'t': [datetime(2000, 1, 1)]}) Out[87]: <xarray.Dataset> Dimensions: (t: 1) Coordinates: * t (t) datetime64[ns] 2000-01-01 Data variables: *empty*
xray now has support (including serialization to netCDF) for
TimedeltaIndex.datetime.timedeltaobjects are thus accordingly cast totimedelta64[ns]objects when appropriate.Masked arrays are now properly coerced to use
NaNas a sentinel value (GH259).
Enhancements¶
Due to popular demand, we have added experimental attribute style access as a shortcut for dataset variables, coordinates and attributes:
In [88]: ds = xray.Dataset({'tmin': ([], 25, {'units': 'celcius'})}) In [89]: ds.tmin.units Out[89]: 'celcius'
Tab-completion for these variables should work in editors such as IPython. However, setting variables or attributes in this fashion is not yet supported because there are some unresolved ambiguities (GH300).
You can now use a dictionary for indexing with labeled dimensions. This provides a safe way to do assignment with labeled dimensions:
In [90]: array = xray.DataArray(np.zeros(5), dims=['x']) In [91]: array[dict(x=slice(3))] = 1 In [92]: array Out[92]: <xarray.DataArray (x: 5)> array([ 1., 1., 1., 0., 0.]) Coordinates: * x (x) int64 0 1 2 3 4
Non-index coordinates can now be faithfully written to and restored from netCDF files. This is done according to CF conventions when possible by using the
coordinatesattribute on a data variable. When not possible, xray defines a globalcoordinatesattribute.Preliminary support for converting
xray.DataArrayobjects to and from CDATcdms2variables.We sped up any operation that involves creating a new Dataset or DataArray (e.g., indexing, aggregation, arithmetic) by a factor of 30 to 50%. The full speed up requires cyordereddict to be installed.
Bug fixes¶
Future plans¶
- I am contemplating switching to the terms “coordinate variables” and “data
variables” instead of the (currently used) “coordinates” and “variables”,
following their use in CF Conventions (GH293). This would mostly
have implications for the documentation, but I would also change the
Datasetattributevarstodata. - I no longer certain that automatic label alignment for arithmetic would be a good idea for xray – it is a feature from pandas that I have not missed (GH186).
- The main API breakage that I do anticipate in the next release is finally
making all aggregation operations skip missing values by default
(GH130). I’m pretty sick of writing
ds.reduce(np.nanmean, 'time'). - The next version of xray (0.4) will remove deprecated features and aliases whose use currently raises a warning.
If you have opinions about any of these anticipated changes, I would love to hear them – please add a note to any of the referenced GitHub issues.
v0.3.1 (22 October, 2014)¶
This is mostly a bug-fix release to make xray compatible with the latest release of pandas (v0.15).
We added several features to better support working with missing values and exporting xray objects to pandas. We also reorganized the internal API for serializing and deserializing datasets, but this change should be almost entirely transparent to users.
Other than breaking the experimental DataStore API, there should be no backwards incompatible changes.
New features¶
- Added
count()anddropna()methods, copied from pandas, for working with missing values (GH247, GH58). - Added
DataArray.to_pandasfor converting a data array into the pandas object with the same dimensionality (1D to Series, 2D to DataFrame, etc.) (GH255). - Support for reading gzipped netCDF3 files (GH239).
- Reduced memory usage when writing netCDF files (GH251).
- ‘missing_value’ is now supported as an alias for the ‘_FillValue’ attribute on netCDF variables (GH245).
- Trivial indexes, equivalent to
range(n)wherenis the length of the dimension, are no longer written to disk (GH245).
Bug fixes¶
- Compatibility fixes for pandas v0.15 (GH262).
- Fixes for display and indexing of
NaT(not-a-time) (GH238, GH240) - Fix slicing by label was an argument is a data array (GH250).
- Test data is now shipped with the source distribution (GH253).
- Ensure order does not matter when doing arithmetic with scalar data arrays (GH254).
- Order of dimensions preserved with
DataArray.to_dataframe(GH260).
v0.3 (21 September 2014)¶
New features¶
- Revamped coordinates: “coordinates” now refer to all arrays that are not used to index a dimension. Coordinates are intended to allow for keeping track of arrays of metadata that describe the grid on which the points in “variable” arrays lie. They are preserved (when unambiguous) even though mathematical operations.
- Dataset math
Datasetobjects now support all arithmetic operations directly. Dataset-array operations map across all dataset variables; dataset-dataset operations act on each pair of variables with the same name. - GroupBy math: This provides a convenient shortcut for normalizing by the average value of a group.
- The dataset
__repr__method has been entirely overhauled; dataset objects now show their values when printed. - You can now index a dataset with a list of variables to return a new dataset:
ds[['foo', 'bar']].
Backwards incompatible changes¶
Dataset.__eq__andDataset.__ne__are now element-wise operations instead of comparing all values to obtain a single boolean. Use the methodequals()instead.
Deprecations¶
Dataset.noncoordsis deprecated: useDataset.varsinstead.Dataset.select_varsdeprecated: index aDatasetwith a list of variable names instead.DataArray.select_varsandDataArray.drop_varsdeprecated: usereset_coords()instead.
v0.2 (14 August 2014)¶
This is major release that includes some new features and quite a few bug fixes. Here are the highlights:
- There is now a direct constructor for
DataArrayobjects, which makes it possible to create a DataArray without using a Dataset. This is highlighted in the refreshed tutorial. - You can perform aggregation operations like
meandirectly onDatasetobjects, thanks to Joe Hamman. These aggregation methods also worked on grouped datasets. - xray now works on Python 2.6, thanks to Anna Kuznetsova.
- A number of methods and attributes were given more sensible (usually shorter)
names:
labeled->sel,indexed->isel,select->select_vars,unselect->drop_vars,dimensions->dims,coordinates->coords,attributes->attrs. - New
load_data()andclose()methods for datasets facilitate lower level of control of data loaded from disk.
v0.1.1 (20 May 2014)¶
xray 0.1.1 is a bug-fix release that includes changes that should be almost entirely backwards compatible with v0.1:
- Python 3 support (GH53)
- Required numpy version relaxed to 1.7 (GH129)
- Return numpy.datetime64 arrays for non-standard calendars (GH126)
- Support for opening datasets associated with NetCDF4 groups (GH127)
- Bug-fixes for concatenating datetime arrays (GH134)
Special thanks to new contributors Thomas Kluyver, Joe Hamman and Alistair Miles.
v0.1 (2 May 2014)¶
Initial release.